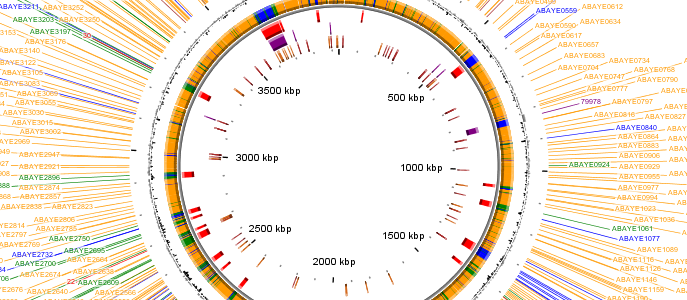
Engineer for the development of new methods in pangenomics
Prokaryotes —bacteria and archaea— are diverse, ubiquitous organisms with vast impacts on health, soil, and ocean ecosystems. Large-scale genome sequencing and pangenomics have revealed their molecular diversity, especially the role of Mobile Genetic Elements (MGEs). Pangenomics analyzes genetic variability across