Background
Today, thanks to the development of metagenomics, our knowledge of microbial diversity has greatly changed. Whereas previously cellular organisms were seen more as independent entities, today we know that they organize themselves into complex microbial communities made up of several species of bacteria, archaea, and eukaryotes and their mobilome (viruses, plasmids and other mobile genetic elements). The proposed project will make it possible to understand the determinants of these complex microbial communities (symbioses/holobionts) through approaches for reconstructing metabolic networks at the scale of each organism but also at the community scale, with a focus on marine ecosystem communities.
Project
This project will be carried out according to 3 complementary axes
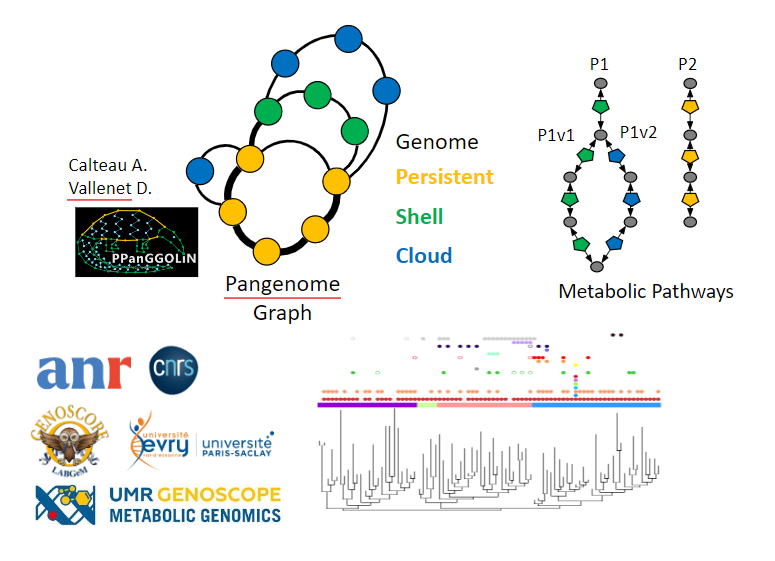
1- Metabolic networks at the scale of genomes and pangenomes.
This first part will aim to reconstruct the metabolic networks at the scale of the genomes of microorganisms but also of the pangenomes of species for which we have several tens or even thousands of genomes. Accessing microbial metabolism from genomic information thus makes it possible to better understand the role of microorganisms, which are major players in biogeochemical cycles and recycling (C, N, S, H), particularly in surface marine and deep oceanic ecosystems.
2- Diversity and evolution of metabolic pathways at the scale of the tree of life
This second part will aim to study the diversity and evolution of metabolic pathways at the scale of the tree of life. We will thus be able to distinguish evolutionary convergences from parallel evolutions and the role of horizontal transfers in the evolution of these metabolic pathways.
3- Metabolic pathways at the ecosystem scale
The purpose of this third part will be to identify and quantify important metabolic pathways for ecosystems and, also, to determine whether the species involved in these metabolic pathways of interest are always the same within similar ecosystems (from different geographical regions).
Linked Projects
ANR Symbiomagnet : Deciphering The Biodiversity, Ecology And Evolution Of Magnetotactic Symbiosis
PhD Project PanGenoThermo : Pangenomic analysis of metabolic pathways in deep ocean living archaea
Publications and presentations:
Presentations and Posters HAL :
Clémence Lauden, Ryan Catchepole, Phil M. Oger, Damien Courtine, Loïs Maignien, et al.. MetaPangenomic analysis within the Thermococcales. EMBO Workshop Molecular biology of Archaea (MBoA) 2024, Jun 2024, Palaiseau, France. 2024. ⟨hal-04707270⟩
Lukáš V F Novák, Marie Hemon, Violette Da Cunha, Karine Alain. Origins and evolution of microbial sulfur disproportionation in the deep sea and beyond. 19th International Symposium on Microbial Ecology ISME19, Aug 2024, Cape Town, South Africa. ⟨hal-05133950⟩
Ryan Catchpole, Clémence Lauden, Damien Courtine, Evelyne Marguet, Stéphanie Fouteau, et al.. Metapangenomic analysis of the metabolic pathway evolution within the Thermococcales. Pangenome 2026 One health, Dec 2025, Valencia (Espagne), Spain. ⟨hal-05504972⟩
Karine Alain, Lukáš Novák, Marilina Fernandez, Stéven Yvenou, Marie Hémon, et al.. Microbial sulfur disproportionation: How much do we know?. 7th international symposium on microbial sulfur metabolisms (7ISMSM), Mar 2026, Lisbonne, Portugal. . ⟨hal-05564516⟩
Publications :
Lukas V F Novak, Lijing Jiang, Marie F Hémon, Marilina Fernandez, Léa Russo, et al.. Sulfur disproportionation occurs globally across anoxic habitats and has multiple mechanisms of independent evolutionary origin. The International Society of Microbiologial Ecology Journal, 2026, 20 (1), pp.wrag042. ⟨10.1093/ismejo/wrag042⟩. ⟨hal-05561923⟩
Financial Support and Institution:
ANR-22-CPJ1-0082-01

