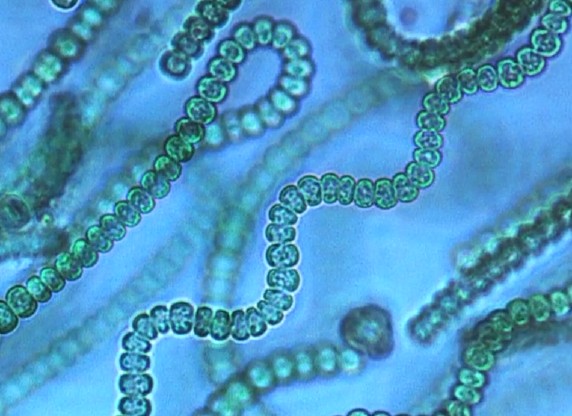
Cyanobacteria are an ancient lineage of photosynthetic bacteria, which are prolific producers of secondary metabolites. Hundreds of bioactive compounds with diverse chemical structures , such as toxins and molecules of pharmaceutical interest, have been characterized from cyanobacteria. In the framework of a collaboration with Muriel Gugger (head of the Pasteur Culture collection of Cyanobacteria – Institut Pasteur), we have shown through a phylum-wide genome investigation, that Cyanobacteria encode a tremendous number of gene clusters devoted to the biosynthesis of secondary metabolites (Shih et al, 2013). Genome mining of secondary metabolites revealed the potential of the Cyanobacteria phylum, to create, diversify and spread the genetic clusters for various compounds: either compounds encoded by NRPS/PKS clusters (Calteau et al, 2014, Pancrace et al., 2017), or ribosomal encoded compounds like cyanobactins (Leikoski et al, 2014). Particularly, we discovered an unexplored potential for siderophore synthesis in cyanobacteria that we will investigate more deeply to understand the different strategies developed in this phylum to capture iron from the environment to maintain their Fe-rich photosynthetic apparatus.
People: Alexandra Calteau
Publications
Pancrace C, Jokela J, Sassoon N, Ganneau C, Desnos-Ollivier M, Wahlsten M, Humisto A, Calteau A, Bay S, Fewer DP, Sivonen K, Gugger M.
Rearranged Biosynthetic Gene Cluster and Synthesis of Hassallidin E in Planktothrix serta PCC 8927.
ACS Chem Biol. 2017 Jul 21;12(7):1796-1804.
Pancrace C, Barny MA, Ueoka R, Calteau A, Scalvenzi T, Pédron J, Barbe V, Piel J, Humbert JF, Gugger M.
Sci Rep. 2017 Jan 24;7:41181.
Pancrace C, Gugger M and Calteau A
Genomics of NRPS/PKS Biosynthetic Gene Clusters in Cyanobacteria
from: Cyanobacteria: Omics and Manipulation (Edited by: Dmitry A. Los). Caister Academic Press, U.K. (2017) Pages: 55-74.
Calteau A, Fewer DP, Latifi A, Coursin T, Laurent T, Jokela J, Kerfeld CA, Sivonen K, Piel J, Gugger M.
Phylum-wide comparative genomics unravel the diversity of secondary metabolism in Cyanobacteria.
BMC Genomics. 2014 Nov 18;15:977.
Leikoski N, Liu L, Jokela J, Wahlsten M, Gugger M, Calteau A, Permi P, Kerfeld CA, Sivonen K, Fewer DP.
Chem Biol. 2013 Aug 22;20(8):1033-43.
Shih PM, Wu D, Latifi A, Axen SD, Fewer DP, Talla E, Calteau A, Cai F, Tandeau de Marsac N, Rippka R, Herdman M, Sivonen K, Coursin T, Laurent T, Goodwin L, Nolan M, Davenport KW, Han CS, Rubin EM, Eisen JA, Woyke T, Gugger M, Kerfeld CA.
Improving the coverage of the cyanobacterial phylum using diversity-driven genomesequencing.
Proc Natl Acad Sci U S A. 2013 Jan 15;110(3):1053-8.
