The MicroScope platform now integrates antimicrobial resistance prediction results generated with AMRFinderPlus 4.2.7, replacing those previously obtained using CARD/RGI, thereby providing up-to-date data under an open license.
AMRFinderPlus uses an AMR protein database, HMMs, a hierarchy of AMR protein families, and a custom rule set to identify AMR genes and point mutations, stress response genes, and virulence genes. The database is highly curated with hierarchical structure for AMR proteins, manually curated cutoffs, and associated hierarchical names. It also includes point mutations, stress response and virulence genes, and it contains additional descriptive fields for each gene or point mutation.
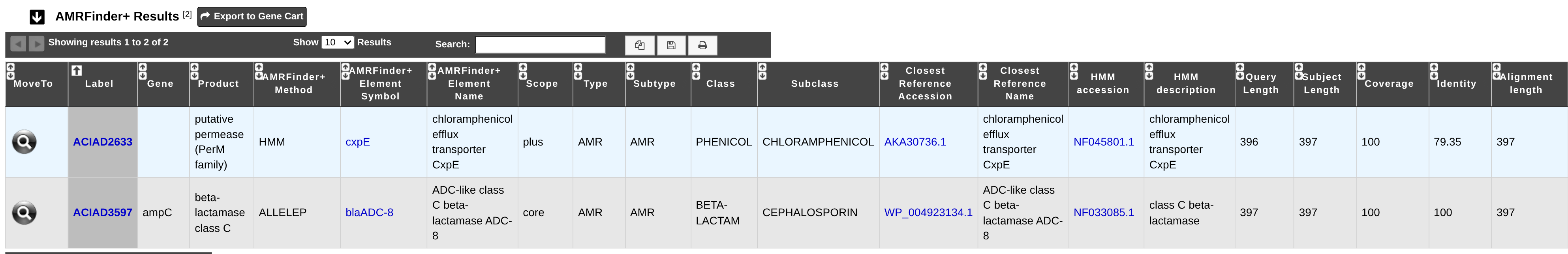
All predictions are available through the Comparative Genomics section, in the main navigation menu and can be searched using the Search by keyword feature. You may also refer to the Resistome section of the MicroScope User documentation for further information.
References: Feldgarden M, Brover V, Gonzalez-Escalona N, Frye JG, Haendiges J, Haft DH, Hoffmann M, Pettengill JB, Prasad AB, Tillman GE, Tyson GH, Klimke W. AMRFinderPlus and the Reference Gene Catalog facilitate examination of the genomic links among antimicrobial resistance, stress response, and virulence. Sci Rep. 2021 Jun 16;11(1):12728. doi: 10.1038/s41598-021-91456-0. PMID: 34135355; PMCID: PMC8208984.
Please report us any bugs you may meet.
